|
1/3/2024 0 Comments Sqlite rowid
The sqlite database is on a very slow machine with tons of disk space, and holds the total archive going back several years. I'm less worried that two corresponding records would contain different values).Ī few more probably irrelevant details: The postgres production system is on a very space-constrained machine, and can only hold about 6 months of data. Is there a better way than counting to validate that records in a specified date range stored in databases on two different systems, one in postgres, one in sqlite, contain the same data? (I presume they do, but I'm a paranoid sort, especially around the idea that my syncing process might somehow drop records.Is there a less janky way to leverage the fact that my data is sorted to get at least some of the benefits of indexing without rolling my own?.If I do decide to insert data out of date order, how can I sort the data and reset the rowids to account for that?.What events would cause the rowid of a given record to change?.Will deleting records create "holes" that sqlite will attempt to "fill in" with subsequent insertions?.Can I therefore implement my own binary search to rapidly find a record with a particular ts? (or use max(rowid) to find the maximum timestamp)?.Is rowid maintained in "insertion order" such that if my data is inserted in order of timestamp, I can assume that a higher rowid will never have a lower timestamp?. 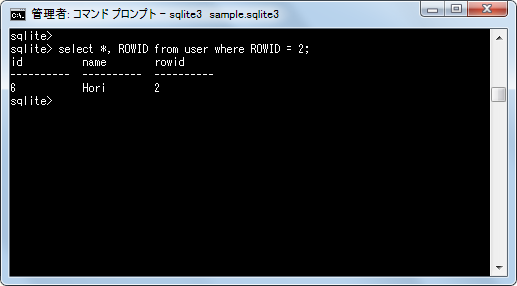
Doing a select count(*) where ts > nnn is slow on the archive (as expected), but it occurs to me that I might be able to use rowid to do a binary search for the first record where ts = nnn and then (perhaps) subtract rowids to get my count of records much faster than the ordinary select would, kind of like a home grown index (or, at very least, allow me to restrict the part of the database that needs to be searched by adding where rowid > mmm and rowid < ooo to the query. To validate that the archive is tracking with my production data, I perform a "checksum" of sorts, counting the number of records in a given date range, and comparing the archive to the production dataset. Multiple records can have the same timestamp, but the timestamp will only increase over time. Therefore, the sqlite db is unindexed.Įvery record has a timestamp (stored in epoch ms), and data is never inserted out of date order. Generally speaking, archive size is more important than archive performance. I am using this as archival backup for working data on a different system (in postgres). This study presents a computational framework for the re-analysis of proteomics datasets to better investigate the viral-host protein interplays upon infection with the Zika virus.I have a large and growing table I'm maintaining in sqlite (250M+ rows). Furthermore, 12 alternative proteins were identified in the proteome profiling of Zika infected monocytes, one of which was significantly up-regulated. Here we show that the use of a customized database including currently non-annotated proteins led to the identification of 4 alternative proteins as interactors of the viral capsid and NS4A proteins. An ever-increasing number of studies have demonstrated the shortcomings of such annotation, which overlooks thousands of functional ORFs. Such databases rely on genome annotations, which enforce a minimal open reading frame (ORF) length criterion. 
Yet these studies used standard human protein sequence databases. In the recent years, several studies have set to identify human host proteins interacting with Zika viral proteins to better understand its pathogenicity. Despite its risk to public health, no antiviral nor vaccine are currently available. Like other flaviviruses, the Zika virus is transmitted by mosquitoes and provokes neurological disorders. The Zika virus is a flavivirus that can cause fulminant outbreaks and lead to Guillain-Barré syndrome, microcephaly and fetal demise.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
